Whole-genome resequencing of S. bicolor x S. halepense Cross Results in Increased Coverage of the Whole Chromosome and Presents New Targets for Future Research
Sorghum, a staple cereal in the semi-arid tropics in Asia and Africa, is becoming popular throughout the world as a healthful, gluten-free alternative to other grains. Introgression with wild lines produces improved cultivars in sorghum as it does in other plants. There has been interest in crosses between Sorghum bicolor and Johnsongrass (S. halepense (L.) Pers.), a wild species, as such breeding can create hybrids with strong perenniality and overwintering. Perennial cultivars are long-lasting cover crops, which through permanently covering the soil, limit its moisture loss and keep the soil biology active, improving its chemical and physical characteristics. Genetically optimizing sorghum lines by introducing these traits is of great interest to breeders and growers alike.
In an effort to work on the introgression of perenniality and overwintering, researchers at the the International Crops Research Institute for the Semi-Arid Tropics in India, in collaboration with Chonnam National University in the Republic of Korea, Sivas University of Science and Technology and Ataturk University in Turkey comprehensively depicted the structure and function of 172 lines. Nineteen of these lines were recombinant inbred lines crossing S. bicolor and S. halepense. The introgression of wild lines into domesticated lines usually results in the introduction of negative traits along with the positive target traits, a situation known as linkage drag. This study used both crossing, backcrossing and selection to minimize the introduction of less desirable traits, such as small-sized kernels, seed shattering, and other wild plant characteristics. The researchers assert that their study is the first to use whole-genome resequencing to “harness” the diversity inherent in the wild S. halepense line. Previous studies used genotyping-by-sequencing, which is limited by low sequencing depth and poor coverage.
The study resulted in a higher density of genes and variants with superior coverage of whole individual chromosomes (>20X) when compared with earlier research studies. Coverage in previous studies resulted in candidate genes and significant SNPs located mostly at distal and proximal ends of the chromosomes. The better variant distribution pattern in the present study opens up the opportunity for boosting gene and major marker discovery in the pericentromeric regions, on which there is little information.
The biological processes GO enrichment analysis presented 5,548 SbxSh private genes which mapped to the GO terms and 34 of these mapped to root system development. Of the 34 root genes, two have been shown to directly effect root growth and development in Arabidopsis thaliana – ROOT PRIMORDIUM DEFECTIVE 1 (RPD1; GeneID: 8054879) and RETARDED ROOT GROWTH (RRG, GeneID: 8072111) (Zhou et al, 2011). The study as a whole shows promise for informing future genomic studies, which can lead to gains in the selected characteristics of domesticated sorghum varieties.
“This work heralded the era of introgressing agroecological relevant traits in agriculturally important crop species; this is in line with the One-Health goal of achieving optimal health for people, animals, plants, and their shared environment.” – Habyarimana
SorghumBase example
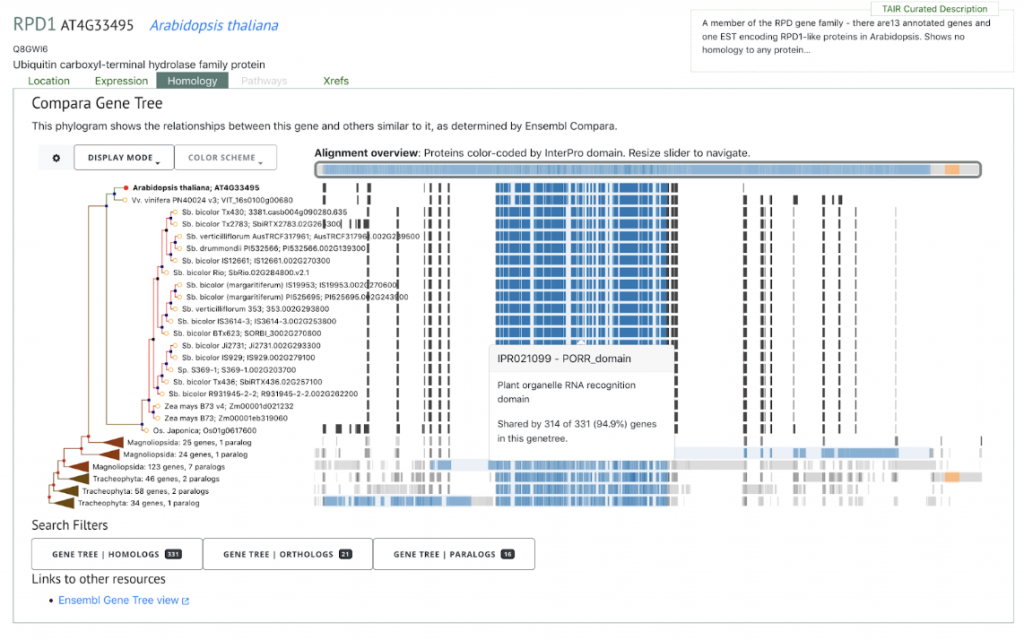
References
Habyarimana E, Gorthy S, Baloch FS, Ercisli S, Chung G. Whole-genome resequencing of Sorghum bicolor and S. bicolor × S. halepense lines provides new insights for improving plant agroecological characteristics. Sci Rep. 2022 Apr 1;12(1):5556. PMID: 35365708. DOI: 10.1038/s41598-022-09433-0. Read more
Zhou X, Li Q, Chen X, Liu J, Zhang Q, Liu Y, Liu K, Xu J. The Arabidopsis RETARDED ROOT GROWTH gene encodes a mitochondria-localized protein that is required for cell division in the root meristem. Plant Physiol. 2011 Dec;157(4):1793-804. PMID: 21984726. DOI:10.1104/pp.111.185827. Read more
Related Project Websites:
https://www.databio.eu/en/


